Projects

Molecular Machines of Poxviruses
Until its eradication the poxvirus Variola caused devastating smallpox epidemics in humans with a fatality rate of 30% and an estimted total death toll of 0.5 billion (!) people. The benign Vaccinia strain was the key to eradicate smallpox and to demonstrate the effectiveness of vaccination campaigns. Still, animal reservoirs of other poxviruses are reponsible for various zoonoses, in particular, sporadic but deadly monkeypox infections. Monkeypox is regarded as a serious threat to cause a future pandemic. Conversely, the non-pathogenic Vaccinia virus has great medical benefit and potential and is currently used as a tool in viral anti-cancer therapies.
Poxviruses are remarkably large and contain a double stranded DNA genome that is replicated and transcribed exclusevely in the cytoplasm of the host cell. Consequently, the virus encodes a complete set of specific transcription factors and a multi-subunit DNA-dependent RNA polymerase. The understanding of the poxviral infection cycle and the underlying molecular biology is therefore of major importance to basic and applied biomedical research.
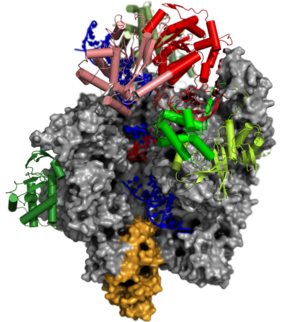
We have recently solved the first structures of the poxviral DNA-dependent RNA polymerase (vRNAP) that facilitates the transcription of all viral genes by single particle cryo-electron microscopy (cryo-EM). The highlights of our poxvirus research are:
- A 0.9 MDa putative packaging complex of the core vRNAP associated with several transcription and processing factors and a cellular tRNA, that we termed complete vRNAP,
- A pre-initiation complex containing the early transcription factor VETF bound to the promoter in the initially melted state,
- A late pre-initiation complex in which a phosphorylated tail of a transcription factor mimics the template strand to keep the complex in an initiation-ready condition, and
- A promoter-escape complex containing a helicase transcription factor that, under ATP consumption, scrunches upstream DNA into the active cleft to generate an energy-loaded intermediate.
Based on these results we were able to devise a mechanistic model of the initial phase of poxviral transcription that, in several aspects, differs excitingly from nuclear transcription mechanisms.
Press Release
Read more ...
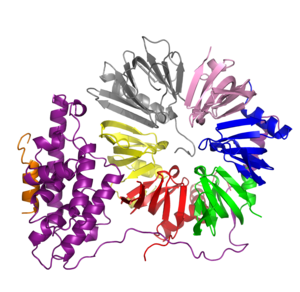
Structure of the SMN-complex
Before genetic information can be translated into proteins non-coding regions, the so called introns, have to be removed from the pre-messenger RNA. The latter is the primary transcript of genetic information contained on the chromosome. Within the eukaryotic cell, this task is facilitated by a huge and complex machinery, the spliceosome. It is not surprising that the assembly of such a complex machine requires, in turn, another complicated machinery. Our research focus is on this molecular assembly machine, the SMN complex. The SMN complex coordinates the formation of basic spliceosomal building blocks called snRNPs. At the heart of this complex is the name-giving survival motor neuron (SMN) protein. Mutations within the gene coding for SMN have been identified as the cause for spinal muscular atrophy (SMA).
We have solved the crystal structures of two key intermediates of the assisted snRNP assembly process. Based on the two protein structures we propose a mechanism how snRNPs are assembled in the crowded environment of the eukaryotic cytoplasm. Press Release
By applying an integrative structural biology approach, we have recently built a model of the complete yeast SMN complex: Press Release
Read more ...
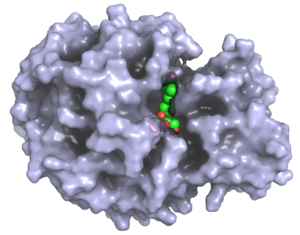
Design of Small Molecule Ligands
The rational design of small, 'drugable' molecules that bind selectively and with high affinity to a particular target protein is a promising field in pharmaceutical and industrial research. In collaboration with the Seibel (department of organic chemistry) and the Sotriffer (department of pharmacy) groups we are developing ligands for enzymes with industrial relevance and for targets as the chaperone Hsc70 and the lectin Galectin-1 that might be useful for anti-tumor therapies.
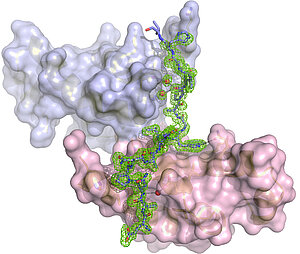
Structural Biology of LARPs
Several La-related (Larp) proteins play an important role in the regulation of the process of translation of messenger RNA into proteins. To understand the molecular details of these regulatory processes, are screening for RNA/protein and protein/peptide complex crystals of Larp proteins.
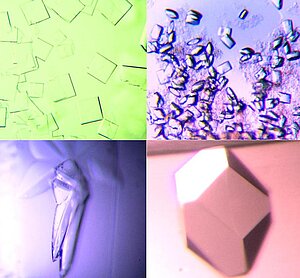

Crystallogenesis
Growing crystals of suitable size and quality is -obviously- the first and most important step of every structure determination project. The ability to increase the success rate during initial crystallization screening as well as crystal optimization will have a significant impact on the overall success rate. Amongst other strategies, we have recently focused on the use of alternative polymeric precipitants to overcome possible limits of the popular polyethylene glycols (PEGs). A commercial screen that exploits the chemical diversity of water soluble polymers as a variable of the 'crystallization space' is marketed as a collaboration with Molecular Dimensions Ltd.