Virus multiplication in 3D
12/12/2019Vaccinia viruses serve as a vaccine against human smallpox and as the basis of new cancer therapies. Two studies now provide fascinating insights into their unusual propagation strategy at the atomic level.
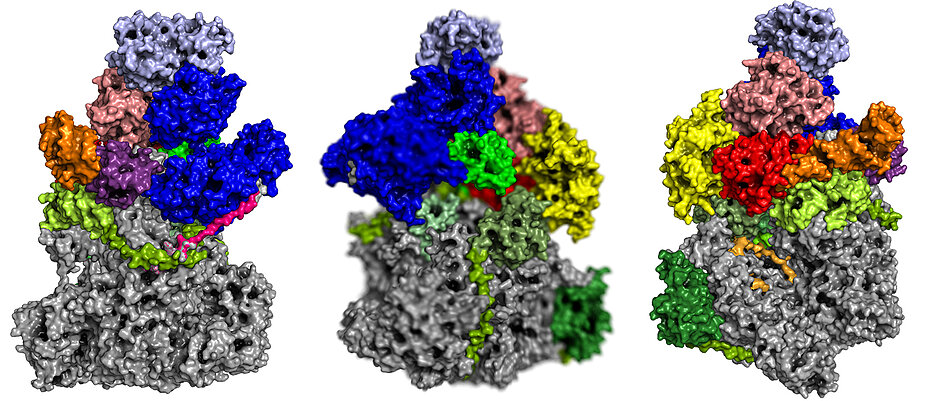
For viruses to multiply, they usually need the support of the cells they infect. In many cases, only in their host’s nucleus can they find the machines, enzymes and building blocks with which they can multiply their genetic material before infecting other cells. But not all viruses find their way into the cell nucleus. Some remain in the cytoplasm and thus must be able to multiply their genetic material independently. To do so, they have to bring their own "machine park". An essential role in this process plays a special enzyme composed of various subunits: the RNA polymerase. This complex reads the genetic information from the genome of the virus and transcribes it into messenger RNA (mRNA), that serves as a blueprint for the proteins encoded in the genome.
Publication in Cell
Scientists from the Biocenter of the Julius-Maximilians-Universität Würzburg (JMU) and the Max Planck Institute (MPI) for Biophysical Chemistry in Göttingen have now succeeded for the first time to solve the three-dimensional structure of a vaccinia virus RNA polymerase at atomic resolution. The vaccinia virus belongs to the poxvirus family, is harmless to humans, and forms the basis of all smallpox vaccines. Due to its benign properties it is currently tested for oncolytic virotherapy, a novel strategy in the combat against cancer.
Responsible for this work are Utz Fischer, Chair of the Department of Biochemistry I of the JMU Würzburg, and Patrick Cramer, Director and head of the Department of Molecular Biology at the MPI for Biophysical Chemistry. In two simultaneous publications in the journal “Cell”, they now present the results of their collaboration.
A molecular clamp that holds everything together
"The vaccinia virus RNA polymerase exists essentially in two forms: the actual core enzyme and an even larger complex, which, thanks to various additional subunits, has special functionalities," explains Fischer. The core enzyme is largely similar to another known enzyme, which has long been the focus of Cramer's department: the cellular RNA polymerase II. This is found in the cell’s nucleus, where it reads the information on the genome and transcribes it into mRNA. Fischer calls the second complex of vaccinia RNA polymerase an ‘all-rounder’. Composed of numerous subunits, it carries out the entire transcription process for the virus, thereby enabling an important step in the pathogen’s multiplication.
The complex is held together by a molecule that the virus lends from its host cell: a so-called transfer RNA (tRNA). This type of molecule normally does not play a role in transcription, but provides the amino acid building blocks for protein production. "Without the involvement of the host tRNA, this huge machinery with all its specific subunits would fall apart," says structural biologist Clemens Grimm, who carried out the structural analysis together with Hauke Hillen of the MPI. The researchers suspect that the tRNA molecule, in addition to its connecting function, performs another important task. "This tRNA can only be loaded with glutamine, an amino acid necessary not only for the production of proteins but also an important source of energy and nitrogen for the cell," explains Aladar Szalay, co-author of the study and director of the Cancer Therapy Research Center (CTRC) at the JMU. Since the virus relies on nitrogen for its replication, the tRNA could serve as a sensor that provides the virus with information about the current nitrogen content in the host cell. If the nitrogen level falls below a certain value, this could be the signal for the virus to leave its host as soon as possible. But this is so far just a hypothesis.
To find out how the viral RNA polymerase works, the researchers also determined its three-dimensional structure during different transcription steps. With these new findings, it is now possible to understand the entire process of virus multiplication on a structural basis. Like in a film, it is possible to track how this molecular machine functions at the atomic level and how the individual processes are choreographed. "What's amazing is how the building blocks of the machine rearrange themselves after the start of transcription to drive the synthesis of the RNA product – this complex is really very dynamic," Hillen explains. To gain this insight, biochemists and structural biologists had to work closely together: The biochemists Julia Bartuli and Kristina Bedenk at the JMU have purified and biochemically characterized the polymerase complex with all its interacting components in a year-long process. The structural biologists Grimm and Hillen were then responsible for determining the three-dimensional structures.
A super microscope provides the necessary data
The researchers obtained the data from a device that has revolutionized structural analysis in recent years: the latest generation cryo-electron microscopes, which are in operation at both the JMU and the MPI. With 300,000 volts, it shoots electrons through samples cooled down to minus 180 degrees Celsius and thus delivers images with a resolution that ranges in the order of atoms. The microscope makes it possible to study biological molecules and complexes and to reconstruct their three-dimensional structure.
For about six months, Grimm and Hillen had to work with their computers until they had developed a spatial model of the polymerase complexes from several terabytes of data. "Without the new cryo-microscopes at our institutions and the excellent cooperation between the two groups, that would not have been possible so quickly and in this quality," says Grimm. With 3D glasses, everyone can now spatially visualize the complex, turn it arbitrarily and dissect it into its subunits.
Among other things, the new findings now offer the possibility to develop inhibitors and modulators to influence the viral propagation cycle. Because Vaccinia replication occurs in the cytoplasm, the scientists also expect it to have therapeutic potential. Currently, studies are underway throughout the world in which Vaccinia viruses are used in the fight against cancer. The company Genelux, which also participated in the study, has already shown in animal experiments and patients the potential of specially optimized Vaccinia viruses to shrink tumors and to detect the smallest metastases. In addition, the researchers expect new and exciting insights into the functioning of the related non-viral RNA polymerase complexes.
A video showing the enzyme complex in detail can be seen on JMU's Youtube channel.
The original publications
„Structural basis of poxvirus transcription: Vaccinia RNA polymerase complexes“, C. Grimm, H.S. Hillen, K. Bedenk, J. Bartuli, S.Neyer, Q. Zhang, A. Hüttenhofer, M. Erlach, C. Dienemann, A. Schlosser, H. Urlaub, B. Böttcher, A. Szalay, P. Cramer and U. Fischer. Cell 179 (2019) pp., 1537-1550, DOI: 10.1016/j.cell.2019.11.024
„Structural basis of poxvirus transcription: transcribing and capping Vaccinia complexes“, H.S. Hillen, J. Bartuli, C. Grimm, C. Dienemann, K. Bedenk, A. Szalay, U. Fischer and P. Cramer: Cell 179 (2019) pp., 1525-1536, DOI: 10.1016/j.cell.2019.11.023
Contact
JMU: Prof. Dr. Utz Fischer, Chair of Biochemistry, Julius-Maximilians-Universität Würzburg, Phone: +49 931 31-84029, utz.fischer@biozentrum.uni-wuerzburg.de
MPI-BPC: Prof. Dr. Patrick Cramer, Department of Molecular Biology, Max Planck Institute for Biophysical Chemistry, Phone: +49 551 201-2800, pcramer@mpibpc.mpg.de