Research
Understanding MYC function
MYC proteins strongly stimulate cell growth and proliferation and alter cellular metabolism to provide building blocks for growth. They also change the interaction of tumor cells with their environment and this includes changes in the interaction of cells with the immune system. Biochemically, they are transcription factors, which and can both activate and repress transcription. We have shown that transcriptional repression takes place in complex with a partner protein, MIZ1, and that repression by this complex is important for MYC-dependent tumor formation.
Surprisingly, MYC proteins essentially bind to all active promoters and to many enhancers in a cell, suggesting that they have a universal function in transcription. Ongoing work in the laboratory aims to understand this function.
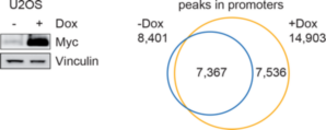
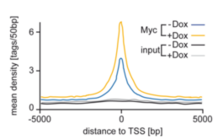
Global binding of MYC to core promoters. Panels on the left show immunoblots from an inducible cell line and a summary of ChIP-sequencing experiments documenting binding of MYC to thousands of promoters. The panel on the right is a metagene analysis of all bound promoters showing that MYC binding sites in promoters are very close to transcription start sites.
Key References
Walz, S., Lorenzin, F., Morton, J., Wiese, K.E., von Eyss, B., Herold, S., Rycak, L., Dumay-Odelot, H., Karim, S., Bartkuhn, M., Roels, F., Wüstefeld, T., Fischer, M., Teichmann, M., Zender, L., Wei, C.L., Sansom, O., Wolf, E., Eilers, M. (2014) Activation and repression by oncogenic Myc shape tumour-specific gene expression profiles, Nature 511:483-7. doi: 10.1038/nature13473.
Wiese, K.E., Haikala, H.M., von Eyss, B., Wolf, E., Esnault, C., Rosenwald, A., Treisman, R., Klefstrom, J., and Eilers, M. (2015) Repression of SRF target genes is critical for Myc-dependent apoptosis of epithelial cells. EMBO J. 34:1554-71.
Vo, B.T., Wolf, E., Kawauchi, D., Gebhardt, A., Rehg, J.E., Finkelstein, D., Walz, S., Murphy, B.L., Youn, Y.H., Han, Y.G., Eilers, M., and Roussel, M.F. (2016) The interaction of Myc with Miz1 defines medulloblastoma subgroup identity. Cancer Cell, 29:5-16.
von Eyss, B., Jaenicke, L.A., Kortlever, R.M., Royla, N., Wiese, K.E., Letschert, L., McDuffus, L.A., Sauer, M., Rosenwald, A., Evan, G.I., Kempa, S., and Eilers, M. (2015) A MYC-driven change in mitochondrial dynamics limits YAP/TAZ function in mammary epithelial cells and breast cancer. Cancer Cell, 28:743-57.
Targeting MYC for tumor therapy
Amplification of MYCN, the gene that encodes N-Myc, is a driver of neuroendocrine tumors such as childhood neuroblastoma and neuroendocrine prostate carcinoma. Like c-Myc, N-Myc is rapidly turned over by the proteasome system and needs to be protected from degradation to be oncogenic. Surprisingly, N-Myc depends on association with the Aurora-A kinase for stabilization. The complex can be disrupted by small molecule ligands of Aurora-A (MLN8237;Alisertib) and these induce degradation of N-Myc. Clinical trials show that Alisertib has therapeutic efficacy in human neuroblastoma and we work on improving therapeutic strategies. Collaborative work with Lars Zender shows that MYC, like N-MYC, is stabilized by complex formation with Aurora-A in hepatocellular carcinoma.
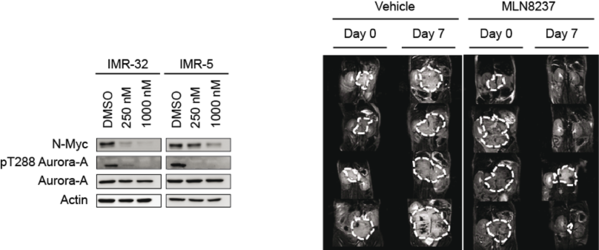
Aurora-A is required to stabilize N-Myc in neuroblastoma. The left panel shows immunoblots of human neuroblastoma cells treated with the Aurora-A kinase inhibitor, Alisertib (MLN8237). Right panel shows the response of mouse neuroblastomas to the drug. Pictures are taken from Brockmann et al., 2013.
Key References
Brockmann, M., Poon, E., Berry, T., Carstensen, A., Deubzer, H.E., Rycak, L., Jamin, Y., Thway, K., Robinson, S.P., Roels, F., Witt, O. Fischer, M., Chesler, L. and Eilers, M. (2013) Small Molecule Inhibitors of Aurora-A Induce Proteasomal Degradation of N-Myc in Childhood Neuroblastoma, Cancer Cell, 24, 75-89.
Dauch, D., Rudalska, R., Cossa, G., Nault, J.G., Kang, T.W., Wuestefeld, T., Hohmeyer, A., Imbeaud, S., Yevsa, T., Hoenicke, L., Pantsar, T., Bozko, P., Malek, N.P., Longerich, T., Laufer, S., Poso, A., Zucman-Rossi, J., Eilers, M., and Zender, L. (2016) A MYC-Aurka protein complex represents an actionable target in p53 altered liver cancer. Nature Medicine, 22(7):744-53. doi: 10.1038/nm.4107.
Wiegering, A., Uthe, F.W., Jamieson, T., Ruoss, Y., Hüttenrauch, M., Küspert, M., Pfann, C., Nixon C., Herold, S., Walz, S., Taranets, L., Germer, C.T., Rosenwald, A., Sansom, O.J., and Eilers, M. (2015) Targeting translation initiation bypasses signaling crosstalk mechanisms that maintain high MYC levels in colorectal cancer. Cancer Discovery, 5:768-81.
MYC and Ubiquitin
Myc proteins and their partners are highly unstable and continuously turned over by the proteasome system. Myc is degraded upon ubiquitination by the Fbxw7 ubiquitin ligase; we found that the ubiquitin-specific protease, Usp28, antagonizes degradation by Fbxw7 both in culture, and in collaboration with Axel Behrens group (LRI) in vivo. Turnover of MYC is also required for transcriptional activation, and MYC associates stably with several ubiquitin ligases that it needs to be active. Together with Nikita Popov we have shown that ubiquitin-mediated turnover of MYC removes inactive and repressive complexes from core promoters.
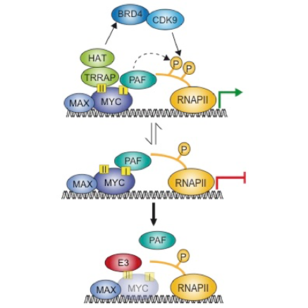
MYC proteins are degraded upon ubiquitination by several ligases. Some of them are required for transcriptional activation by MYC and we suggest that these ligases remove repressive complexes of MYC from core promoters (see Peter et al. and Jaenicke et al. 2015).
Key References
Jaenicke, L.A., von Eyss, B., Carstensen, A., Wolf, E., Xu, W., Greifenberg, A., Geyer, M., Eilers, M., and Popov, N. (2015). Ubiquitin-dependent turnover of MYC antagonizes MYC/PAF1C complex accumulation to drive transcriptional elongation. Molecular Cell, 61:54-67.
Peter, S., Bultinck, J., Myant, K., Jaenicke, L.A., Walz, S., Müller, J., Gmachl, M., Treu, M., Boehmelt, G., Ade, C.P., Schmitz, W., Wiegering, A., Otto, C., Popov, N., Sansom, O., Kraut, N., and Eilers, M. (2014) Tumor cell-specific inhibition of MYC function using small molecule inhibitors of the HUWE1 ubiquitin ligase. EMBO Molecular Medicine 6:1525-41.
Popov, N., Schülein, C., Jaenicke, L. and Eilers, M. (2010) Ubiquitination of the Myc amino-terminus by beta-TrCP antagonizes Fbw7-mediated degradation. Nature Cell Biology, 12,973-81.