Projects
Assembly of RNA Protein complexes
(see Fischer et al., 2011; Chari and Fischer, 2010; Neuenkirchen et al., 2008)
In this research focus we aim to understand how RNA-protein complexes (RNPs) are assembled within the context of a living cell. To address this question, we use the small nuclear ribonucleoprotein (snRNP) particles of the spliceosome as a model system. We have identified a complex network of assembly factors (“assembly chaperones”) that guide and facilitate RNP formation. Our current focus is the functional and structural investigation of the assembly machinery.
We combine biochemical and structural methods, as well as experimental strategies in living systems (cells and model organisms), to understand how RNA-protein complexes (termed RNPs) assemble in vivo to form a functional unit and how they act in their designated cellular pathway. Our main object of study are RNPs involved in the maturation of eukaryotic messenger RNAs (mRNAs), which are synthesized as precursors that need to be extensively processed. The major processing step on almost all mRNAs of higher eukaryotes is splicing, in which the non-coding introns are removed and coding exons are precisely joined to generate the open reading frame, the blueprint for ribosomal translation into protein.
Splicing is catalyzed by the spliceosome, which is probably the most complex machine within our cells. It is composed of the small nuclear ribonucleoproteins (snRNPs) U1, U2, U4/6 and U5 and probably more than 100 additional proteins. These units assemble anew on each intron of an mRNA facilitate removal of introns.
The cell has developed intricate strategies to generate snRNPs and the spliceosome itself. This process involves nucleo-cytoplasmic transport events and the action of sophisticated molecular machines that regulate and catalyze the assembly reaction.
U snRNP assembly: Formation of U snRNPs starts with the transport of newly transcribed snRNA to the cytoplasm. A set of common (Sm) proteins is then loaded onto U snRNAs, leading to the assembly of a ring-shaped Sm core domain. In a next step, the monomethyl-guanosine cap of the snRNA is converted to a trimethyl-guanosine cap. Eventually, the assembled snRNP particle is imported into the nucleus, where also specific proteins are added to complete the biogenesis pathway. We have recently shown that the assembly of U snRNPs requires assisting factors that are united in the SMN complex, consisting of the SMN protein and eight additional proteins termed Gemin2-8 and unrip. This macromolecular unit recruits Sm proteins as well as U snRNAs and mediates the formation of the Sm core domain. An additional unit termed PRMT5 complex cooperates with the SMN-complex in snRNP assembly. The PRMT5 complex functions as an assembly chaperone, by enabling the formation of Sm protein higher order structures that are required for their subsequent transfer onto the SMN-complex. The detailed molecular and structural analysis of the assembly machinery (i.e. the SMN-complex and the PRMT5-complex) is an ongoing project in this research area.
(see Chari et al., 2008; Meister and Fischer, 2002; Meister et al., 2001; Meister et al., 2001; Kroiss et al., 2008)
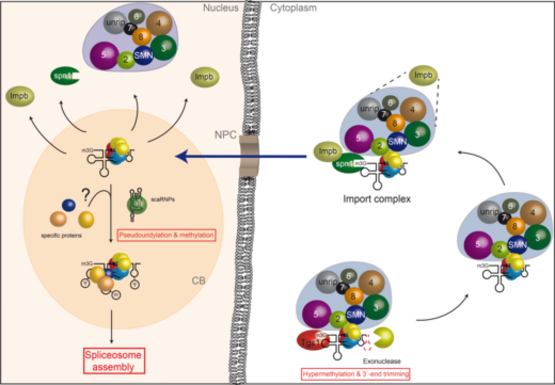
After 3´-end trimming by a exo-ribonuclease and hypermethylation of the m7G-cap by Tgs1, Spn1 associated with Importin binds the 2,2,7-trimethylguanosine (m3G) cap structure. The Sm core domain represents a second import signal. The SMN-complex and/or a yet unknown additional factor allow binding to Importin. After nuclear import and localization in Cajal Bodies the import complex dissociates. SnRNP specific proteins join the core snRNP and nucleotides within the snRNA are modified by scaRNPs. Mature snRNPs can now engage with the splicing machinery.
Spliceosome assembly: Once U snRNPs have been assembled and imported to the nucleus, they can integrate into the spliceosome. Spliceosome formation likewise requires specific proteins. We could show a role of the SMN-homologous protein SMNrp (also termed SPF30), in the integration of the tri-snRNP into the pre-spliceosome. We are hence persuing the possibility that spliceosome formation requires assembly factors that are functionally and/or structurally related to components of the SMN-complex.
After each splicing reaction, the spliceosome must be dis-assembled to allow fromation of a new spliceosome on another intron that needs to be spliced. In addition, snRNPs undergo a steady turnover and hence mechanisms must exist that allow for the degradation of „used“ snRNPs. Understanding these „recycling“ processes is another focus of our group.
Assembly of U7 snRNP: In contrast to spliceosomal snRNPs, the U7 snRNP is not invovled in pre-mRNA splicing but rather in the processing of 3’-ends of histone mRNAs. This particle contains a unique set of proteins and hence requires a specialized chaperone system for its formation. In collaboration with the group of Daniel Schümperli (University of Bern) we have shown that a specialized SMN-complex mediates assembly of U7 snRNP (see Pillai et al., 2003). This assembly machinery is currently investigated by biochemical and structural means.
Post-translational modifications and the regulation of RNP assembly
Studies by several groups including our own have uncovered that the assembly process of snRNPs is tightly controlled by means of post-translational modifications. This allows the integration of snRNP biogenesis with the metabolism of the cell. Arginine- methylation of some Sm proteins of snRNPs appears to be a key regulatory event in the early phase of assembly (see Meister et al., 2001). We have identified the methyltransferase responsible for this modification and currently analyze its kinetics and role in snRNP formation. Several proteins of the SMN-complex are modified by phosphorylation and this appears to regulate the activity and sub-cellular localization of the assembly machinery. These modifications and their role in snRNP biogenesis are analyzed in a collaboration with Oliver Gruss (ZMBH Heidelberg).