Group Dr. Daniel Maag
Dr. Daniel Maag
Lehrstuhl für Pharmazeutische Biologie
Julius-von-Sachs-Institut für Biowissenschaften
Julius-von-Sachs-Platz 2
97082 Würzburg
Phone: + 49 931 31-85538
Fax: + 49 931 31-86182
daniel.maag@uni-wuerzburg.de
CV
Publications
Research focus
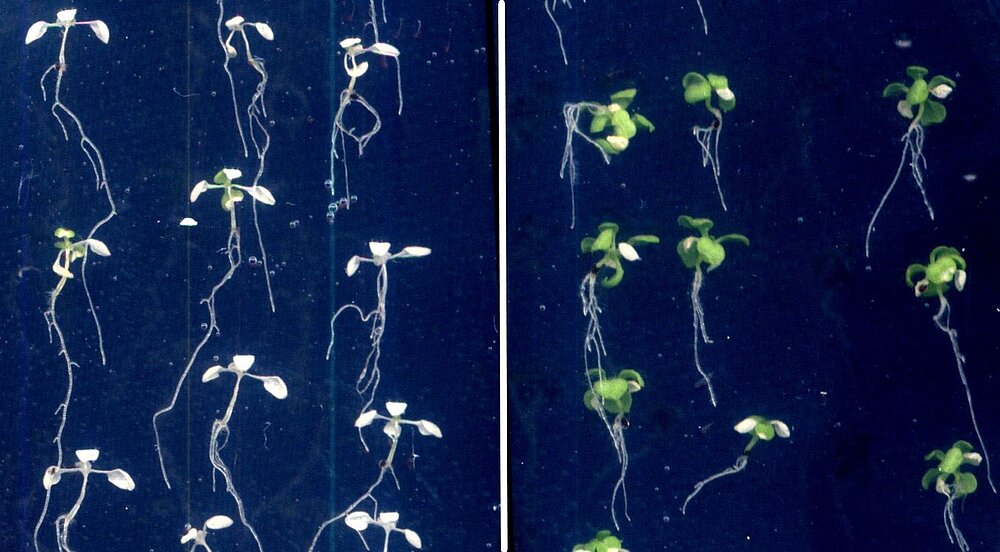
In their native habitat, most plants face a constantly changing temperature environment potentially threatening their reproductive success. However, plants are able to sense these temperature changes and adapt their growth and development correspondingly. In my group, we study different aspects of plant responses to elevated temperatures using the model organism Arabidopsis thaliana ranging from very rapid responses at the molecular level to morphological adjustments over the course of several days. Ultimately, our research aims at a better understanding of temperature perception by plants as well as the underlying regulatory networks, which will be a prerequisite for the breeding of climate change-resilient crop varieties.
Early temperature signalling during thermomorphogenesis
Upon moderate heat stress, Arabidopsis seedlings display a growth response syndrome summarised as thermomorphogenesis, which results in an increased leaf cooling capacity.
In this project, we aim at the identification of molecular regulators that are involved in the early signalling underlying thermomorphogenic growth responses using warm temperature-induced hypocotyl elongation as a model response.

Identification of novel regulators of the heat shock response
Recent studies suggest that within A. thaliana considerable natural variation exists concerning the extent of the thermal response. In this project, we make use of this intraspecific variation to identify novel regulators of the heat shock response by using an unbiased genome mapping approach that takes advantage of the available Arabidopsis genetic resources. Recently, the genomes of more than 1000 A. thaliana accessions have been sequenced and made publically available by the 1001 Genomes Consortium. This large collection of accessions spanning A. thaliana’s natural geographic range serves us as an excellent resource for studying the genetic basis of acquired thermotolerance using genome-wide association studies (GWAS). This project is performed in close collaboration with the Korte Group at the University’s Centre for Computational and Theoretical Biology (CCTB).