Research
Research topic 1: The oncogenic function of MYC
Almost all human tumors express significantly more MYC protein than healthy tissue, and mouse models have shown a clear relationship between MYC activity and the development of cancer. Thus, although MYC is a crucial oncogene, for many years MYC was not considered a promising therapeutic target because the MYC protein is also essential for healthy cells. With the motivation to find out whether and how the oncogenic function of MYC differs from its physiological mode of action, I have intensively investigated the mechanism of MYC-mediated gene regulation in recent years and been able to make two fundamental observations.
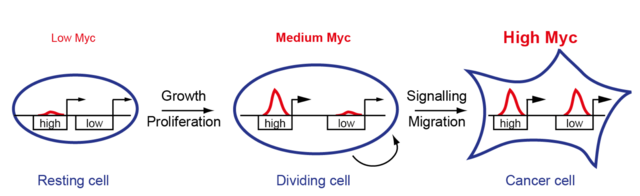
First, we were able to show that MYC is a universal transcription factor that binds to the promoters of all expressed genes and can regulate them (Walz et al, Nature, 2014). Secondly, we found that the target genes differ significantly in their affinity for MYC (Lorenzin et al, eLife, 2016). High affinity target genes are regulated at low MYC concentrations, as is the case in healthy cells, and are important for cell growth and proliferation. At very high MYC levels, as found in tumor cells, MYC also binds to low affinity promoters and this binding leads to the regulation of genes that mediate tumor-specific processes (Vo*, Wolf*, et al, Cancer Cell, 2016). In the future, we want to identify the target genes of MYC that mediate its oncogenic properties and attack them pharmaceutically.
Research topic 2: Investigation of transcriptional elongation
While the target genes of MYC have been extensively analyzed by other research groups and us in recent years, the actual mechanism by which MYC regulates target genes was unclear. It was also unclear whether MYC fundamentally alters the transcriptional process of cells, as seems possible considering the global binding behavior of MYC. In order to investigate how and why MYC alters the transcription machinery globally, my research group has set up two experimental systems: we both carried out a series of mass spectrometric investigations with MYC and the RNA polymerase and analyzed the global transcriptional behavior (4sU-sequencing, ChIP-sequencing).
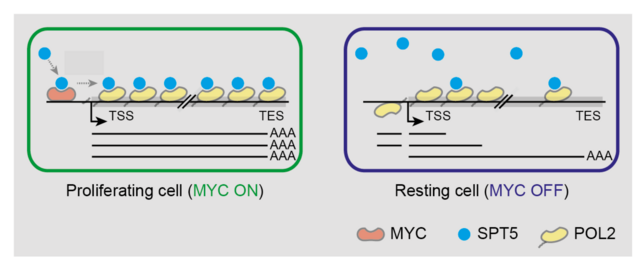
These data result in the surprising and consequential model that, contrary to expectations, MYC does not primarily change the initiation (=transcription rate), but instead loads the RNA polymerase with elongation factors. If this loading mechanism is absent when MYC is inactivated, the RNA polymerase loses a great deal of its processing power. Therefore, the inactivation of MYC leads to the production of abortive, non-protein-coding transcripts instead of intact mRNAs (Baluapuri et al, Molecular Cell, 2019). This observation raises the question of what the biological role of this abortive transcription is.
Research topic 3: New pharmaceutical concepts in oncology
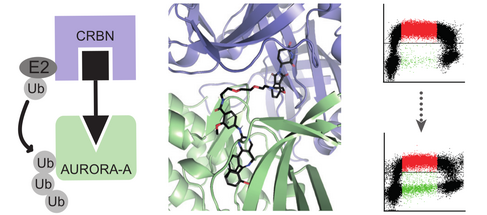
While the results of our scientific work primarily establish or modify fundamental models of the mode of action of MYC and the course of transcription, they also provide opportunities for the development of new pharmaceutical concepts. Thus, the identification of tumor-relevant target genes as well as of interaction partners of MYC offer new therapeutic targets. However, many interaction partners of MYC, as well as MYC itself, are difficult drug targets, as they often lack an enzymatic mechanism of action. We, therefore, investigated whether the new substance class of "PROTACs" can be used to reduce the concentration of MYC and its interaction partners in tumors in a therapeutically relevant way. A PROTAC directed against the MYC-interacting Aurora-A protein has already shown very promising properties.