Model interpretability in medical image segmentation
Summary
The state of the art in automatic image segmentation is currently reached through deep learning models. These models proof to have good performance on the data sets they are trained and evaluated on. However, it is often hard to estimate how well these models perform on novel datasets or if inputs have different characteristics than those seen during training. Sensitivity analysis is a systematic approach to analyze and quantify the effect that certain perturbations on the input have on the output. For example we can visualize the effect that rotation has on the segmentation performance of a cardiac MRI image. In addition to visualizing the effect qualitatively we can also quantify the performance using a metric like the DICE-score.
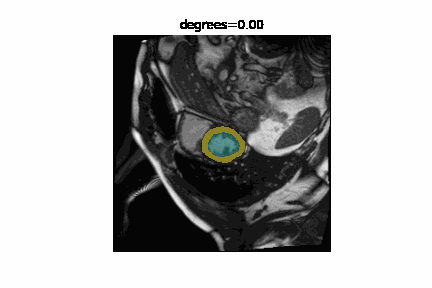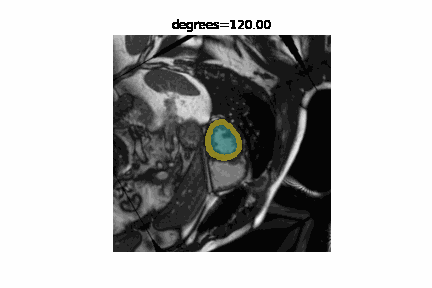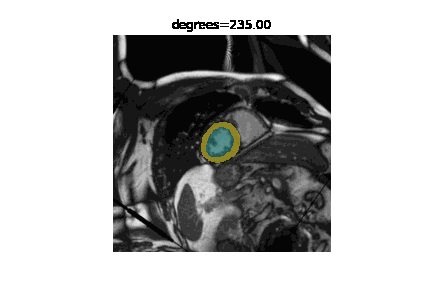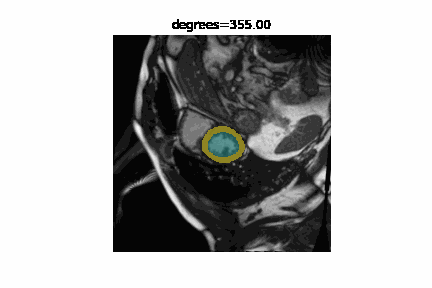
We developed misas a software that helps to easily perform sensitivity analysis on arbitrary segmentation models with a range of perturbations (e.g. rotation, cropping, zooming, introducing MR artifacts, ...).
While this software is published [1] and in a stable state, we have plans to further develop the method. Our planned work includes:
- automatically generating reports for a single or multiple inputs
- adding the capability to compare different models on the same data set
- updating the underlying packages
- add more case-studies demonstrating the utility of this method for real research questions
[1] Ankenbrand, M. J., Shainberg, L., Hock, M., Lohr, D., & Schreiber, L. M. Sensitivity analysis for interpretation of machine learning based segmentation models in cardiac MRI. BMC Medical Imaging, 21(27). https://doi.org/10.1186/s12880-021-00551-1