Priv.-Doz. Dr. Ulrich Terpitz | |
Telefon: | +49 931 31-84226 |
| Since 2011 | Lecturer and Groupleader, Department of Biotechnology & Biophysics(Biocenter), University of Würzburg, Germany |
| 2009-2011 | Postdoctoral Researcher, Max-Planck-Institute of Biophysics, Frankfurt am Main, Germany |
| 2005-2009 | PhD in Biochemistry, Johann-Wolfgang-Goethe University and Max-Planck-Institute of Biophysics, Frankfurt am Main, Germany |
| 2004 | Research Assistant, University of Morelos, Cuernavaca, Mexico |
| 2001-2004 | Study of Biology, Friedrich-Schiller-University Jena, Germany |
| 1998-2001 | Study of Biology, University of Hamburg, Germany |
We combine super-resolution microscopy and electrophysiology (Patch-clamp technique) to analyse proteins that are crucial for the interaction of microbes with their environment. Within the collaborative research centre TR124 FungiNet (project A2) we use state-to-the-art fluorescence microscopy for unravelling the interaction between innate immune cells and Aspergillus fumigatus. Due to their tiny size and their often-strong autofluorescence, fluorescence microscopy of fungi is challenging. We therefore recently established - and currently optimize - protocols for the visualisation of fungal hyphae by expansion microscopy (ExM), structured illumination microscopy (SIM), direct stochastic optical reconstruction microscopy (dSTORM), and correlative light and electron microscopy (CLEM). In our electrophysiological studies, we focus on light-gated membrane proteins, especially on fungal rhodopsins. Although these green-light sensing proteins are widespread in the fungal kingdom, still little is known about their physiological function in fungal life cycle. Fungal rhodopsins also provide the fundament for developing and characterising optogenetical tools that allow for analysis of physiological downstream processes upon light-trigger in isolated cells.
Our group is part of the collaborative research center CRC/TR 124 FungiNet and involved in project A2. This collaborative research centre combines the expertise of researchers in Jena and Würzburg and aims to decipher common principles of fungal pathogenesis and infection in the interplay with the innate immunity and adaptive immune system. A better understanding of fungal diseases is of fundamental interest, as the number of life-threatening infections due to opportunistic fungal pathogens has drastically increased over the past two decades, whereas the clinical diagnosis of these often-mortal infections is still very difficult.
Within A2, we complement classical immunological and molecular approaches by super-resolution microscopy (Fig. 1) to unravel A. fumigatus-host myeloid/lymphoid cell interactions at the cellular and molecular level. We investigate fundamental processes involved in the immune response of natural killer (NK) cells against Aspergillus fumigatus. We are specially interested in NK-cell receptors interacting with the fungus, with special focus on the newly described A. fumigatus receptor CD56 and corresponding pathogen-associated molecular patterns on the fungus.
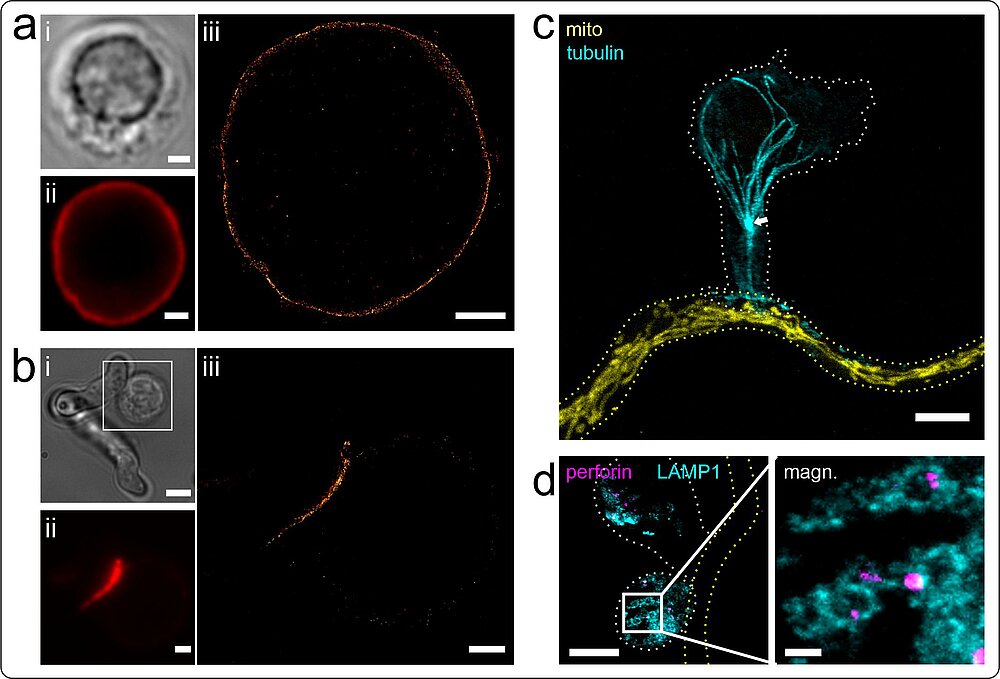
Ziegler et al., Sci. Rep. 7, 6238 (2017), Trinks et al., Commun. Biol. 4, 1151 (2021)
Fungi use photoreceptors to adapt to changing light conditions in their environment. Green light is sensed by fungal rhodopsins that belong to the type I (microbial) rhodopsins. Microbial rhodopsins are widespread in the fungal kingdom and consist of seven transmembrane helices forming an interior pocket for the chromophore all-trans retinal, which is covalently bound to the protein via a protonated Schiff-base (Fig. 2). Upon light-activation fungal rhodopsins act as proton pumps or sensory proteins. We analyse the pump function of distinct fungal rhodopsins from different filamentous fungi including the ascomycetes Fusarium fujikuroi and Aureobasidium pullulans as well as the basidiomycete Ustilago maydis.
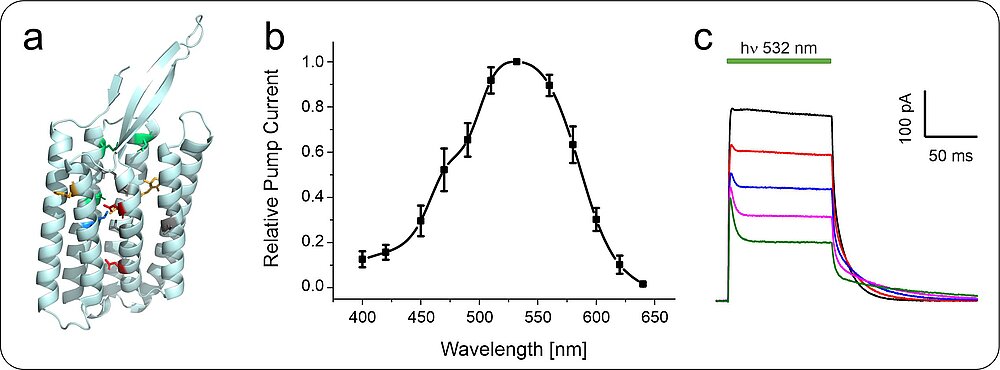
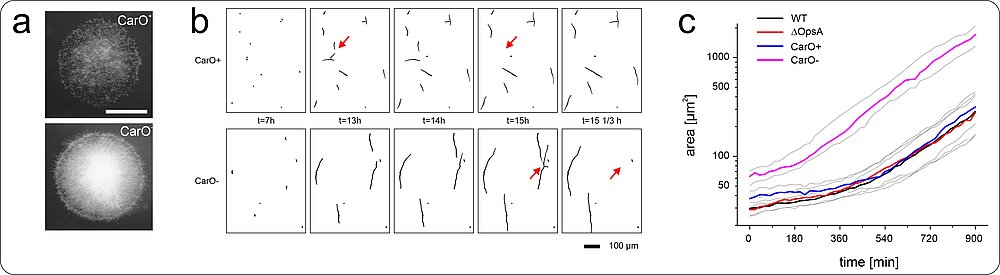
In general, detailed knowledge regarding the physiological function and biological role of fungal rhodopsin is still missing. As many substantial processes in filamentous fungi such as reproduction and pathogenicity are controlled by light, we aim to unravel the task of these green light sensors in fungi. Our recent data from the rhodopsin CarO of the rice pathogen F. fujikuroi suggest a role of rhodopsins in conidia germination (Fig. 3) and the fungus-plant interaction.
Fungal rhodopsins are also of interest for optogenetic applications as some of them like ApOps2 and ApOps3 from A. pullulans are very strong proton pumps that can be converted to proton channels by site-directed mutagenesis.
García-Martínez et al., Sci. Rep.5, 7798 (2015); Adam et al., Int. J. Mol. Sci., 19, 215 (2018); Panzer et al., Front. Microbiol. 10, 735 (2019); Panzer et al., Front. Mol. Biosci., 8:750528 (2021).
We developed an implementation for the open source software ImageJ called “HyphaTracker”, allowing for computer-assisted analysis of fungal area alteration over time. This plugin is optimized for the analysis of germinating spores and allows for automatic area analysis of the fungal germlings after manual image processing. HyphaTracker allows for analysing the germination profile of the conidia of rhodopsin-deficient F. fujikuroi strains versus their reference strains and by that for revealing differences in the dynamics of the conidia germination. Before performing area analysis, the application filters the recordings and excludes particles and crossing hyphae from the analysis (Fig. 3).
The imageJ toolbox “HyphaTracker” can be downloaded here.
Brunk et al., Sci. Rep. 8, 605 (2018)
Adam, A., Deimel, S., Pardo-Medina, J., García-Martínez, J., Konte, T., Limón, M., Avalos, J., Terpitz, U.: Protein activity of the Fusarium fujikuroi rhodopsins CarO and OpsA and their relation to fungus–plant interaction. Int. J. of Mol. Sci. 19, 215 (2018)
Brunk, M., Sputh, S., Doose, S., van de Linde, S., Terpitz, U.: HyphaTracker: An ImageJ toolbox for time-resolved analysis of spore germination in filamentous fungi. Sci. Rep. 8, 605 (2018).
García-Martínez, J., Brunk, M., Avalos, J., and Terpitz, U., The CarO rhodopsin of the fungus Fusarium fujikuroi is a light-driven proton pump that retards spore germination. Sci. Rep. 5, 7798 (2015)
Panzer, S., Brych, A., Batschauer, A., Terpitz, U., Opsin 1 and opsin 2 of the corn smut fungus Ustilago maydis are green-light-driven proton pumps. Front. Microbiol. 10, 735 (2019).
Panzer, S., Zhang, C., Konte, T., Bräuer, C., Diemar, A., Yogendran, P., Yu-Strzelczyk, J., Nagel, G., Gao, S., Terpitz, U., Modified rhodopsins from Aureobasidium pullulans excel with very high proton-transport rates. Front. Mol. Biosci. 8, 750528 (2021).
Trinks, N., Reinhard, S., Drobny, M., Heilig, L., Löffler, J., Sauer, M., Terpitz, U., Subdiffraction-resolution fluorescence imaging of immunological synapse formation between NK cells and A. fumigatus by expansion microscopy. Commun. Biol. 4, 1151 (2021).
Ziegler, S., Weiss, E., Schmitt, A.-L., Schlegel, J., Burgert, A., Terpitz, U., Sauer, M., Moretta, L., Sivori, S., Leonhardt, I., Kurzai, O., Einsele, H., Löffler, J., CD56 is a pathogen recognition receptor on human natural killer cells. Sci. Rep. 7, 6138 (2017)