Light-activated Cyclases: PAC & Cyclop
Light-activated Cyclases: PAC & Cyclop
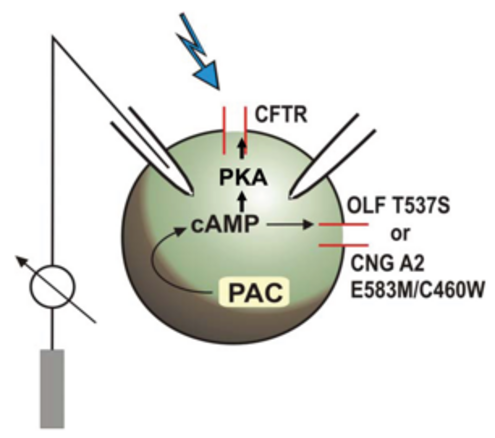
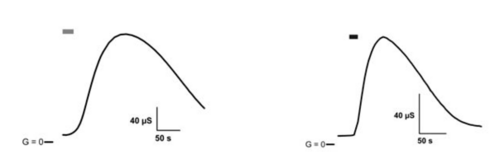
Already in 2002, a Japanese working group discovered light-activated adenylyl cyclases in the monocellular flagellate Euglena gracilis (Iseki et al., 2007, Nature) which they named photoactivated adenylyl cyclase (PAC) a and PAC b. The first functional heterologous expression and use of PAC from Euglena (EuPAC) as an „optogenetic tool“ in cultured animal cells and the living fruit fly Drosophila melanogaster (by Martin Schwärzel, now FU Berlin) was shown by us, in collaboration with other groups, in 2007 (Schröder-Lang et al., Nature Methods), see fig. 1.
Other PACs were found in genome data from microbes and were characterized by us and others. Particularly attractive is PAC from the bacterium Beggiatoa spec. (bPAC), since it is only 350 amino acids in size and has a high ratio of activity in light to activity in the dark (L / D). By mutagenesis and gene fusion we are currently changing and improving the properties of bPAC.
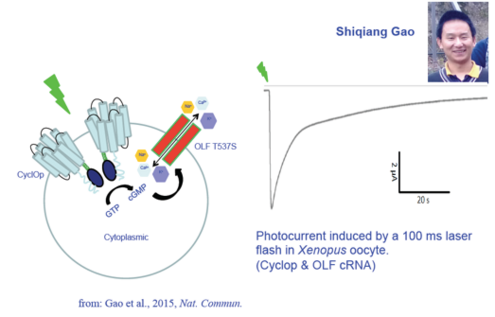
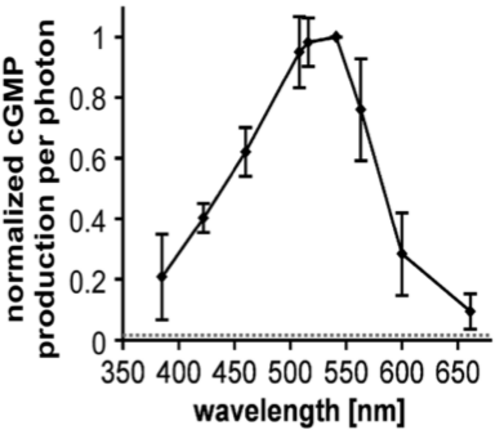
In 2014, a Brazilian group showed that a rhodopsin fusion with a guanylyl cyclase is expressed in the aquatic fungus Blastocladiella emersonii and is activated by green light (Avelar et al., 2014, Curr. Biol.). We synthesized the DNA and characterized the protein after heterologous expression in oocytes or HEK293 cells. After extensive characterization (Gao et al., 2015, Nat. Commun.), we named the protein Cyclase Opsin or Cyclop, and also showed light-activated inward currents after coexpression of a cGMP-sensitive cation channel, see Fig. 2.
We mutated Cyclop proteins from different organisms to produce an adenylyl cyclase from the guanylyl cyclase: Cyclop-PAC. The light-activated cAMP increase was shown by us in-vitro with membrane fragments from oocytes and in-vivo in neurons by the group of Dr. Christine Gee (ZMNH, Hamburg). This work was published together with the structure of the soluble cyclase part of Cyclop, as obtained by the group of Peter Hegemann (Scheib et al., 2018).