Convergent evolution of carnivorous plants
Convergent evolution of carnivorous plants
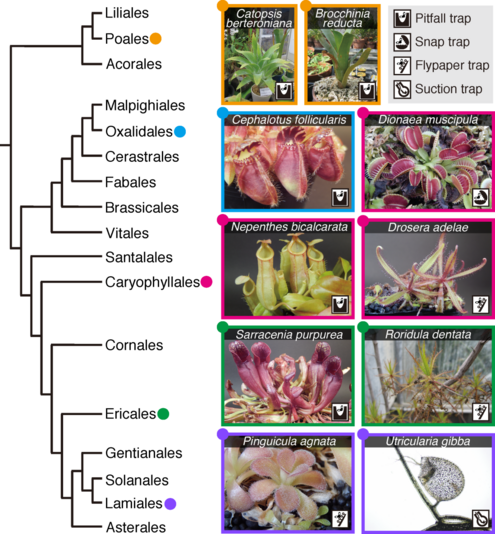
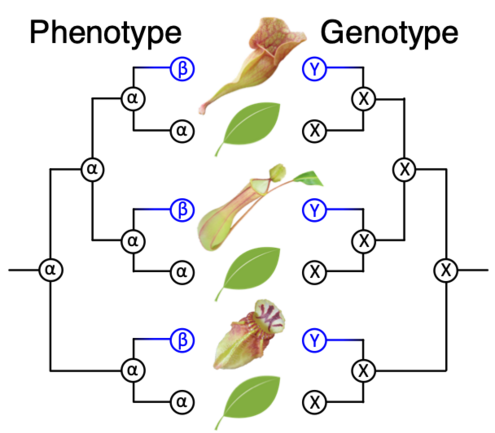
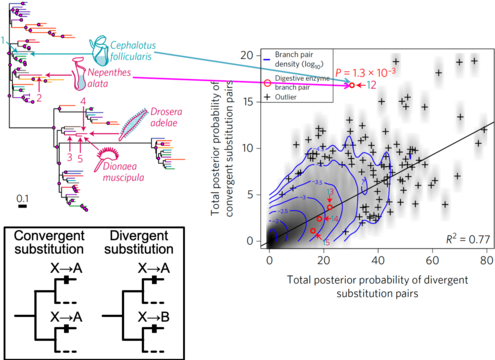
The repetitive nature of convergent evolution opens a venue to link genetic changes with phenotypic changes that arose during the evolution of complex traits in multicellular lineages. An excellent example of such convergent evolution is the trait of carnivory, which arose in five independent orders of flowering plants. Because all carnivorous plants form specialized leaves capable of attracting, trapping, and digesting prey and absorbing nutrients, they represent a classical example of a set of organisms with convergently evolved complex traits. To understand the link between convergent genes and convergent phenotypes in the complex trait evolution of carnivorous plants, we will apply novel bioinformatics and molecular genetic techniques that enable genome-wide detection of molecular convergence associated with phenotypes. The proposed research builds on our central hypothesis that common genetic changes underlie the evolution of carnivory-related traits, a hypothesis that is strongly supported by our finding of molecular convergence in multiple digestive enzymes (Fukushima et al., 2017). Multiple carnivorous lineages will be analyzed to detect convergent changes in gene repertoires, gene expression, and protein sequences that occurred concurrently with the emergence of carnivory, with particular attention to gene duplication events. Convergent genes will be further characterized in planta mainly using Cephalotus follicularis, an emerging model plant for the study of carnivory. We will use CRISPR-based genetic modifications to revert convergent genetic changes and measure the impact to carnivorous traits and associated protein properties to analyze the phenotypes under ancestral genotypes. We expect that this project will facilitate our understanding of how complex traits evolved and how biological systems respond to natural selection in a predictable manner during macroevolution.