Congenital hearing loss
Lab Members:
Dr. Michaela A.H. Hofrichter, PhD
Julia Doll, PhD student
Tabea Röder, Technician
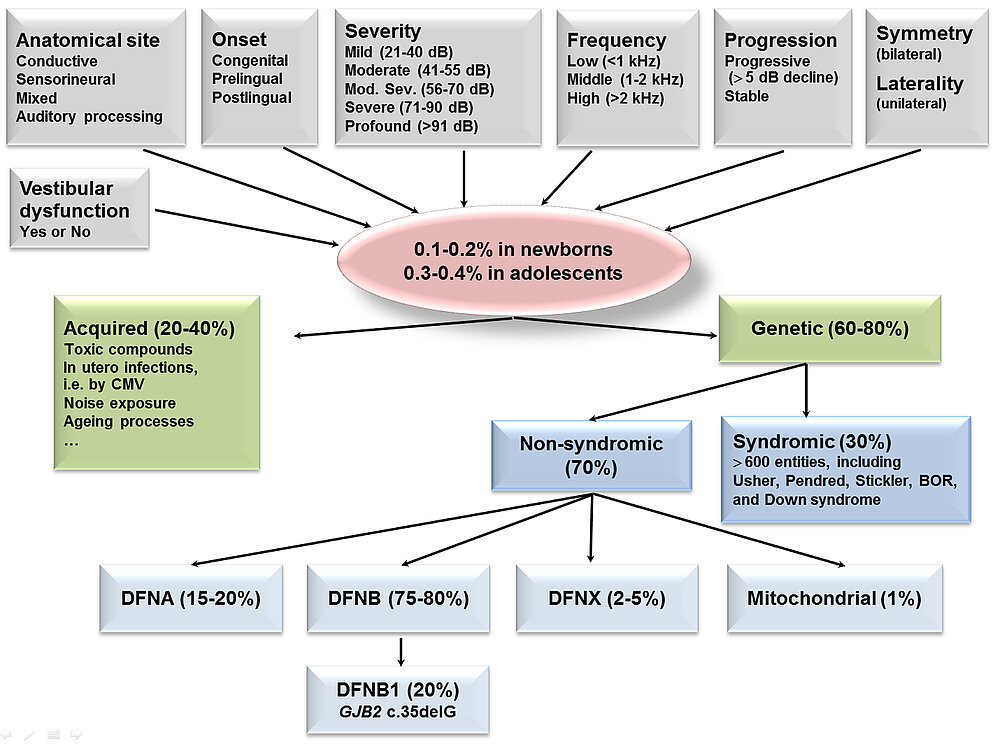
Congenital sensorineural hearing loss (HL) is among the most common birth defects, occurring in 1-2 out of 1,000 newborns. Congenital HL often leads to impaired development of speech and language, as well as poor performance in school and loss of cognitive potential. It may also impair the individual’s social and emotional relationships. Among known causes, genetic and environmental factors play integral roles. In developed countries approximately 80% of congenital HL has a genetic basis. Hearing impairment is extremely heterogeneous. Clinically, it can be further classified by virtue of the affected anatomical site(s), age of onset, severity, affected frequencies, progression, symmetry/laterality, and presence/absence of vestibular dysfunction (Fig. 3). Genetic forms can be divided into non-syndromic (NSHL) and syndromic HL (SHL) with the majority (~70%) of cases representing NSHL. To date, >100 genes have been associated with NSHL including autosomal dominant, autosomal recessive, and X-linked NSHL loci. Notably, one gene, GJB2 accounts for 20-30% of children with autosomal recessive HL in many populations. In addition to NSHL, there are ~600 syndromes including HL as one symptom. The absence of additional symptoms in children does not necessarily exclude SHL. Hearing impairment can be the first manifestation in a long list of clinical features in individuals with SHL. It is estimated that 20-30% of patients diagnosed with NSHL are syndromic cases. The complexity of the auditory system with numerous specialized cell types and systematic mouse screens suggest that up to 1,000 genes are essential for hearing, most of which remain to be discovered. We use state-of-the art next generation sequencing approaches to identify the underlying gene defects in HL patients from different populations.
Hofrichter M.A.H., Doll J., Habibi H., Enayati S., Vahidi Mehrjardi M.Y., Müller T., Dittrich M., Haaf T., Vona B. Exome-wide copy number variation analysis identifies a COL9A1 in frame deletion that is associated with hearing loss. Eur. J. Med. Genet. 62, 103724 (2019).
Hofrichter M.A.H., Mojarad M., Doll J., Grimm C., Eslahi A., Hosseini N.S., Rajati M., Müller T., Dittrich M., Maroofian R., Haaf T., Vona B. A conserved p.Arg108 residue in S1PR2 (DFNB68) is fundamental for proper hearing: evidence from a consanguineous Iranian family. BMC Med. Genet. 19, 81 (2018).
Vona B., Nanda I., Hofrichter M.A.H., Shehata-Dieler W., Haaf T. Non-syndromic hearing loss gene identification: a brief history and glimpse into the future. Mol. Cell. Probes 29, 260-270 (2015).
Vona B., Hofrichter M.A.H., Neuner C., Schröder J., Gehrig A., Hennermann J.B., Kraus F., Shehata-Dieler W., Klopocki E., Nanda I., Haaf T. DFNB16 is a frequent cause of congenital hearing impairment: implementation of STRC mutation analysis in routine diagnostics. Clin. Genet. 87, 49-55 (2015).
Vona B., Müller T., Nanda I., Neuner C., Hofrichter M.A.H., Schröder J., Bartsch O., Läßig A., Keilmann A., Schraven S., Kraus F., Shehata-Dieler W., Haaf T. Targeted next-generation sequencing of deafness genes in hearing-impaired individuals uncovers informative mutations. Genet. Med. 16, 945-953 (2014).
Schneider E., Märker T., Daser A., Frey-Mahn G., Beyer V., Farcas R., Schneider-Rätzke B., Kohlschmidt N., Grossmann B., Bauss K., Napiontek U., Keilmann A., Bartsch O., Zechner U., Wolfrum U., Haaf T. Homozygous disruption of PDZD7 by reciprocal translocation in a consanguineous family: a new member of the Usher syndrome protein interactome causing congenital hearing impairment. Hum. Mol. Genet. 18, 655-666 (2009).
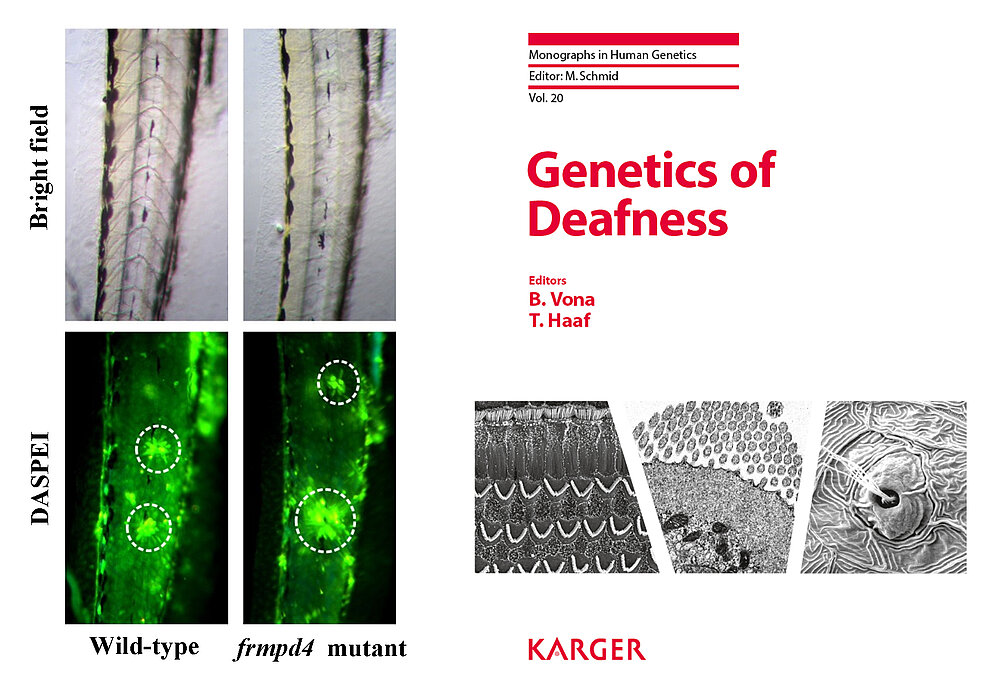
Approximately 70% of all human genes have a zebrafish ortholog. We have established a specialized zebrafish laboratory for functional analyses of newly identified HL genes (Fig. 4). The main advantages of this model are the ease of genetic manipulation, large clutch sizes, inexpensive maintenance and minimal amounts of facility space. The accessibility of the auditory organs permits in vivo imaging of fluorescently labelled neuromasts and scanning electron microscopy. HL mutant lines are characterized by behavioural acoustically evoked response testing.
Vona B., Doll J., Hofrichter M.A.H., Varshney P., Haaf T., Varshney G.K. Small fish, big prospects: using zebrafish to unravel the mechanisms of hereditary hearing loss. Hear. Res. doi10.1016/j.heares.2020.107906 (2020).
Through collaborations we systematically analyse patients/families with unknown syndromic disorders. In particular, consanguineous families can facilitate the identification of new disease genes.
Lekszas C., Foresti O., Raote I., Liedtke D., König E.M., Nanda I., Vona B., De Coster M., Cauwels R., Malhotra V., Haaf T. Biallelic TANGO1 mutations cause a novel syndromal disease due to hampered cellular collagen secretion. Elife 9, e51319 (2020).
Kariminejad A., Szenker-Ravi E., Lekszas C., …, Haaf T., …, Vleminckx K., Nikuei P., Reversade B. Homozygous null TBX4 mutations lead to posterior amelia with pelvic and pulmonary hypoplasia. Am. J. Hum. Genet. 105, 1294-1301 (2019).
Nava C., Dalle C., Rastetter A., …, EuroEPINOMICS RES Consortium, Haaf T., Leguern E., Depienne C: De novo mutations in HCN1 cause early infantile epileptic encephalopathy. Nature Genet. 46, 640-645 (2014).