summer_school_2014

Modern Methods in Infection Biology
3rd International Summer School
September 15th - 19th 2014
Julius-Maximilians-University Würzburg Germany
The workshop provides a major opportunity for networking with local experts in the field, as well as establishing new networks with international peers.
Selection criteria
Outstanding BSc and MSc students, and PhD candidates who have a clear focus on infection biology. Applicants are selected on the basis of motivation and qualification. International and national applications are welcome, as are applications from outstanding students of the University of Würzburg.
The number of participants is limited to 25.
Early registration is recommended.
Course requirements
Advanced spoken and written English skills are required as all lectures and labs are held in English.
Certification/ECTS
Participants will receive a certificate of participation as well as 5 ECTS.
Accommodation
Updated: (April, 2014)Currently rooms in Schönstatt Centre have been booked out over our lecture time, please check the babelfish hostel for accomodation. https://www.babelfish-hostel.de
Limited accommodation for international students is provided at the Schönstatt-Zentrum Marienhöhe, Josef-Kentenich-Weg 1, 97074 Würzburg, 5 minutes walking distance to the Biocentre.
Anfahrt: http://www.schoenstatt-wuerzburg.de/bwo/dcms/sites/bistum/extern/schoenstatt_wuerzburg/anfahrt.html
Altervative:
When Schönstatt Centre is full, there are no more rooms available, the participants could book the rooms from this youth hostel on the webpage: https://www.babelfish-hostel.de
Cultural programme
Welcome Dinner,
Historic City Tour
Farewell Dinner,
Programme organisers
Prof. Dr. Th. Dandekar
Dr. U. Rapp-Galmiche
This programme is funded by the Ministry of Education and Research, grant number 01PL11019.


The University of Würzburg is listed amongst the top four German universities in the Life Sciences and has a strong focus on infection biology research (source: funding ranking DFG).
The Faculty of Biology of the University of Würzburg is organizing an annual international Summer School in infection biology. Speakers and scientific coordinators are Prof. Dr. Dandekar, Prof.
Dr. Brehm, Dr. Kramer and Dr. Fraunholz.
Our goal is to offer national and international BSc, MSc, and PhD students the opportunity to meet, learn and discuss, in order to develop and apply cutting edge insights and techniques and view these from a bioinformatics as well as a microbiology perspective.
Important new methods in drug screening, microscopy and system biology will be discussed and demonstrated. Participants will gain a deeper insight into important models of infectious diseases and will benefit from small group discussions and subsequent application of newly acquired knowledge and techniques.
Application information
The application deadline is to be announced.
Please send applications to:
Dr. Ulrike Rapp-Galmiche
Josef-Martin-Weg, Gebäude 54/1
Campus Hubland-Nord
Universität Würzburg
97074 Würzburg
Germany
Phone: +49-0931-3186901
Email: ulrike.rapp@uni-wuerzburg.de
Applications should include:
-
Letter of motivation (100 words)
-
Curriculum vitae
-
Publications, if applicable.
- A limited number of travel grants are available upon request
Workshop location
Biocentre of the University of Würzburg,
Am Hubland, 97074 Würzburg
Germany
Programme details
Lectures and labs will be held daily from 9.00 a.m. to 5.15 p.m. The main meeting point is lecture theatre A 103.
Monday September 15th
Parasitic Helminths
Prof. Dr. K. Brehm (Homepage)
Medical Parasitology
Institute of Hygiene and Microbiology
Figure: Echinokokkus
On the first day of the workshop the labs and lectures will not be at the Biocentre but at the University Clinics Department Hygiene and Microbiology (Haus E1).
Please find the written directions and maps below:
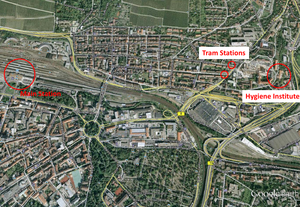
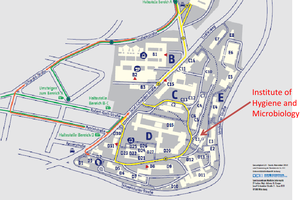

(please click the pictures to enlarge them)
You can reach the main area of the University Hospital with the streetcar routes 1 and 5. You get on the streetcar at the Hautpbahnhof Ost (Main Station East) in the direction of Grombühl and get off at stop “Uniklinikum Bereich D (University Hospital area D; Petrinistrasse)". You can reach the Institute of Hygiene and Microbiology by walk (5 min) as indicated on the map. (The walk from the Main Station takes about 20-25 minutes.)
Parasitic Flatworms: A tale of mortality and immortality.
Parasitic helminths are a major cause of so-called ‘Neglected Diseases’. And Neglected Diseases are called ‘neglected’ because nobody (or only very few) has/have an interest to study them. So, why are we doing it? This is something we’re going to figure out in the ‘Parasitic Helminth’-day of the summer school. We shall learn that the development of novel therapies is not the only motivation to do infectious disease research and that these parasites (like many others) are highly fascinating from the view-point of immunologists as well as cell-, developmental- and evolutionary biologists. Above all, we shall learn how molecular research can be done in a ‘non-mainstream’ setting and which strategies should be followed to establish an infectious disease model system from scratch.
Schedule
9:00-10:30 Introduction: Parasitic helminths and the spellbinding world of tapeworm infections (including tapeworm movie; demonstration of specimen; narration of the ultimate tapeworm joke)
10:30-10:50 Coffee break
10:50-12:00 Interactive Teaching: how to develop a research model system from scratch.
12:00-13:00 Lunch break
13:00-14:20 Würzburg’s current tapeworm research projects: PhD students report
14:20-15:50 Practical aspects of tapeworm and larval tapeworm research (in small groups)
15:50-16:10 Coffee break
16:10-18:00 Final discussion (including Quiz Show ‚Who wants to be a Parasitonaire?‘)
Tuesday September 16th
Systems Biology of Infection
Prof. Dr. Th. Dandekar (Homepage)
Chair, Bioinformatics
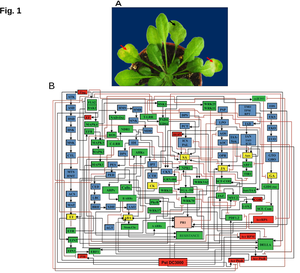
Pseudomonas Syringae attacks the host -- how do you understand the host response?
(A) Phenotype: Pst (104 CFU/ml) infection; red arrows indicate pathogen inoculation; black arrow indicates mock (10mM MgCl2) inoculation. Symptoms weare photographed three days post pathogen inoculation (DPPI). (B) Network: Topology of Pst- mediated hormone disease networks in Arabidopsis. Connectivity among nodes is based either on activation (->) or inhibition (-|). Node designation: blue, enzymes of hormone biosynthesis and degradation; yellow, active hormone molecules; green, host regulatory factors; red, Pst- originated pathogenicity factors responsible for triggering immunity in Arabidopsis; pink, “PR-1”, marker node for immunity against the infection of Pst in Arabidopsis. All nodes are denoted by abbreviations.
More information on these questions is found in our plant cell paper: Naseem M, Philippi N, Hussain A, Wangorsch G, Ahmed N, Dandekar T. Integrated systems view on networking by hormones in Arabidopsis immunity reveals multiple crosstalk for cytokinin. Plant Cell. 2012 May;24(5):1793-814.
However, the same techniques can also be applied (and are shown in the course) to study human infection defense against pathogens.
Schedule
Location: lecture theatre A102 and CIP pool, You find that easily, if you are in the main hall or Foyer of the Biocentre of the University of Würzburg
https://www.biozentrum.uni-wuerzburg.de/en/about-the-biocenter/how-to-get-here//
Begin: 9:00
Equipment: computers – these are provided (CIP pool)
Introduction: Systems biology of infection – bioinformatical approaches
Prof. Thomas Dandekar 9:00-10:00 (lecture theatre A102)
Part I Databanks and Resources 10:00-11:30 (CIP pool in the basement, we will lead you to it if not yet known)
Dr. Chunguang Liang (PlasmoDB, Top targets in Malaria, Protecs database EcoliHub / MetaCyc, Trypanosome DB)
Here is, in addition, the link to PlasmoDB:
http://plasmodb.org/plasmo/
The GO fight project:
http://gofightagainstmalaria.scripps.edu/index.php/project-details
Small, unsuccessful database
http://www.malariajournal.com/content/8/1/178
http://www.bioinformatics.org/project/?group_id=737
The TDR Targets database v5 (a nice one)
http://tdtargets.org/
Part II Genome analysis and annotation 11:30-12:30 (lecture theatre A102)
Paul Costea (EMBL Heidelberg; metagenomics sequencing, relevance for infectious diseases, species variation within a global community, sub-species stratification)
Lunch break 12:30-13:30
Prof. Peer Bork 13:30-14:00 (lecture theatre A102)
Metagenomics of the human gut
Disussion 14:00-14:15
Part III Modelling metabolic networks 14:15-15:00 (CIP pool)
Dr. Alexander Cecil (how to do elementary mode analysis; simple examples on Staphylococci from own research)
Coffee break 15:00-15:30
Part IV Signalling in infectious biology 15:30-17:00 (CIP pool)
Dr. Muhammad Naseem (using Squad, gene expression data, semi-quantitative models, in essence the cell paper)
Part V Current trends in systems biology in infectious biology 17:00-18:00 (lecture theatre A102)
Prof. Thomas Dandekar (we will show and discuss current trends in systems biology with a focus on computer-based approaches, what they can and what they can not elucidate)
Checks and balances of the host against the pathogen better eludicated:
PLOS pathogen paper
Optimize Antibiotic therapy looking at timing and combinations
http://www.nature.com/news/mapping-identifies-best-targets-for-malaria-prevention-1.10781
--> Janina Zirkel Paper
Optimizing Ebola containment and therapy in Africa
Wednesday September 17th
Round table day and discussion and lab visits
Location: Lecture Theatre A103
discussion: future and perspectives of infection biology 9:00-09:45
(participants: all; guests: Prof.s Dandekar, Brehm, Dr. Fraunholz, Dr. Kramer, Dr. Rapp-Galmiche)
Dr. Xaver Kober 10:00-10:30
(Studies: M.Sc., Fokus Master of Life Science, PhD, GSLS excellency school)
Herr Jürgen Seufert 10:45-11:15
(Visiting programs; International Office, accomodation & stay; direct study admission at faculty of biology, English international program)
Thursday September 18th
Intracellular S. aureus
Dr. M. Fraunholz (Homepage)
Microbiology (Lectures: A 103; Practicals: CIP pool)
Figure: Staphylococcus
Topic areas: Intracellular bacterial host-pathogen interactions, microscopy and image analysis demonstrated here by means of Intracellular S. aureus
Upon infection Staphylococcus aureus is readily phagocytosed by macrophages and neutrophils but is also to invade epithelial and endothelial cells, fibroblast, osteoblasts, and keratinocytes. The uptake by latter so-called non-professional phagocytes has been proposed to play a role in evasion of the innate immune system. Further survival of phagolysosomal killing by S. aureus may also lead to disseminating infections within migrating phagocytes. We and others have shown that S. aureus is capable of escaping from host cell phagosomes. One of our methods is microscopy based and relies on the recruitment of a cytoplasmically expressed host cell marker that is recruited to the bacterial cell wall upon phagosomal membrane rupture. Another tool in the laboratory is photoactivatable GFP (PA-GFP), which allows to selectively activate the fluorescence by laser light and thus „label“ as well as follow dynamics/movements of the organisms.

Depicted are two different strains of S. aureus (magenta), of which the left is uanable to escape from phagosomes, whereas the one on the right translocates to the host cell cytoplasm and thereby recruits the escape marker (green). Scale Bar: 5 µm.
On the third day of the summer school we will address theoretical and practical aspects of confocal laser scanning microscopy as well as photoactivatable proteins. We will analyze images using the powerful freeware Fiji/ImageJ and will even introduce macro writing that can be used by the participants in other aspects of research as well.
Location: Lecture Theatre A103
Friday September 19th
Trypanosomes (Lectures, practical: A 103)
Dr. S. Kramer (Homepage)
Zoology I
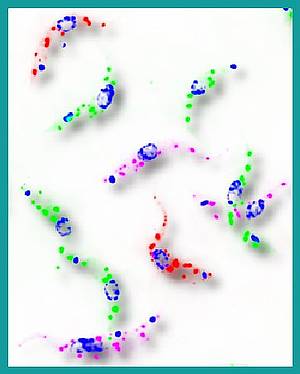
The African Trypanosome: A killer and a survival genius
African trypanosomes cause African Sleeping Sickness as well as the related cattle disease Nagana, affecting people’s health both directly and indirectly in sub-saharan Africa. During the ‘Trypanosome-day’ of the summerschool we will look at the parasites from the point of view of a parasitologist, but also from the point of view of a cell biologist: you will see that trypanosomes are model organisms for many biological questions. A non-conventional teaching method will be used to explore the non-conventional biology of trypanosomes: Pantomine. We will also have a look at the ‘real beast’ by high resolution microcospy and we will get an insight view into the actual research that is currently taking place in the four trypanosome labs that are settled in Würzburg (Engstler, Janzen, Siegel, Kramer).
Schedule
Location: Lecture Theatre A103
9:00-9:45 Introduction lecture to Trypanosomes / Safety instructions
9:45-10:30 Movie (African sleeping sickness)
10:30-11:00 Coffee break
11:00-12:30 Reading time
12:30-13:30 lunch break
13:30-14:30 Current Research in Würzburg trypanosome labs: PhD students report
14:30-16:30 Preparing the pantomime presentation (Coffee break in between) parallel High resolution microscopy of life trypanosomes (in small groups)
16:30-18:00 Pantomine presentations
Course links
![]()

![]()