Abstract
Schwarz R, Liang C, Kaleta C, Kühnel M, Hoffmann E, Kuznetsov S, Hecker M, Griffiths G, Schuster S, Dandekar T.
Integrated network reconstruction, visualization and analysis using YANAsquare. BMC Bioinformatics. 2007 Aug 28;8:313. PubMed PMID: 17725829; PubMed Central PMCID: PMC2020486.BACKGROUND:
Modeling of metabolic networks includes tasks such as network assembly, network overview, calculation of metabolic fluxes and testing the robustness of the network.
RESULTS:
YANAsquare provides a software framework for rapid network assembly (flexible pathway browser with local or remote operation mode), network overview (visualization routine and YANAsquare editor) and network performance analysis (calculation of flux modes as well as target and robustness tests). YANAsquare comes as an easy-to-setup program package in Java. It is fully compatible and integrates the programs YANA (translation of gene expression values into flux distributions, metabolite network dissection) and Metatool (elementary mode calculation). As application examples we set-up and model the phospholipid network in the phagosome and genome-scale metabolic maps of S.aureus, S.epidermidis and S.saprophyticus as well as test their robustness against enzyme impairment.
CONCLUSION:
YANAsquare is an application software for rapid setup, visualization and analysis of small, larger and genome-scale metabolic networks.
YANAsquare
YANAsquare
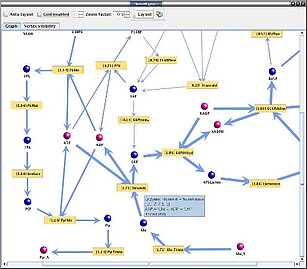
-YANAsquare Software: Download
-YANA Software: Download
-Metatool: Download