QUORUM SENSING
Quorum sensing (Engstler lab)
Parasites must not overwhelm the host
Thus, their population should not get out of control.
Quorum is from the Latin word quōrum = of whom
Many bacteria use a system of quorum sensing to correlate gene expression with cell density.
How does SIF work in the host's circulation?

A cow has 40 liters of blood.
Stumpy induction
About 20 years ago, a quorum sensing pathway was discovered that controls development of the African sleeping sickness pathogen Trypanosoma brucei. The parasites secrete the (still elusive) ‘stumpy induction factor’ (SIF) that drives proliferating slender trypanosomes into the cell cycle arrested stumpy stage. In this way, the parasites limit their cell density and hence, the burden they impose on their mammalian host. It has further been proposed that only SIF-induced stumpy trypanosomes can successfully infect the transmitting insect vector, the tsetse fly.
What is SIF?
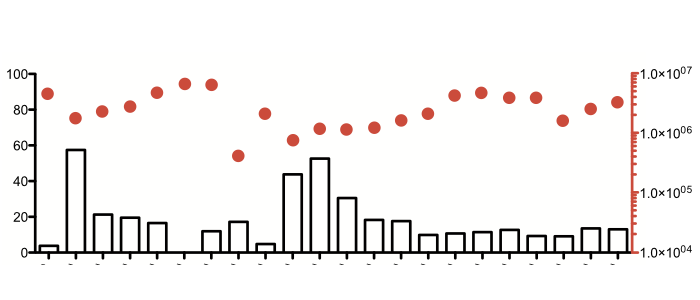
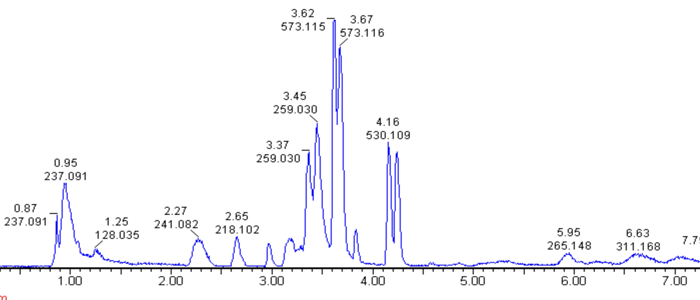
Our work aims at identifying SIF by combining liquid chromatography (LC), untargeted metabolomics and activity bioassays using a pleomorphic reporter cell line. In a first purification step conditioned cell culture medium containing SIF was subjected to methanolic protein precipitation extraction. The resulting deproteinated concentrate was further purified by reversed-phase solid phase extraction eluted with 10% methanol in water, which demonstrated the highly polar nature of SIF. We thus opted for hydrophilic interaction LC using an amide column in order to separate polar metabolites. Surprisingly, the active fractions contained pyruvate as a possible candidate. In fact, pyruvate can mimic SIF action, however, only at concentrations that appear unphysiological. We have data that rule out that pyruvate is SIF, but that also suggest that it might be a related compound.
Publications
We still have not identified SIF, but we hope to have sometime soon. Meanwhile, we have also worked on SIF-independent stumpy induction:
Zimmermann, H., Subota, I.; Batram, C, Kramer, S, Janzen, C, Jones, NG, Engstler, M (2017) A Quorum Sensing-independent Path to Stumpy Development in Trypanosoma brucei. Plos Pathogens doi: 10.1371/journal.ppat.1006324
Batram, C., Jones, N.G., Janzen, C.J., Markert, S.M., and Engstler, M. (2014). Expression site attenuation mechanistically links antigenic variation and development in Trypanosoma brucei. Elife 3, e02324.








