Telomere structure & function (THIS PROJECT WAS DISCONTINUED IN 2024)
The Shelterin Complex
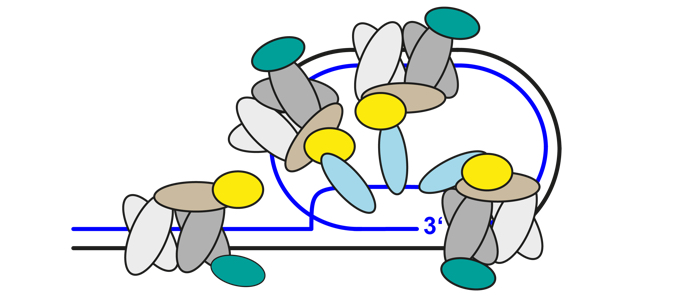
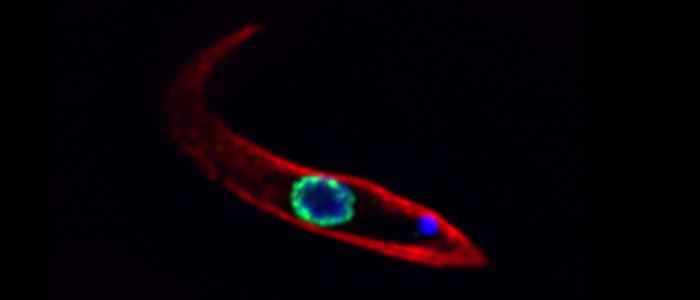
Protects the ends of chromosomes
Structure and function of telomeres
The telomeric ends of eukaryotic chromosomes are protected by nucleoprotein complexes. The telomeric protein complex in mammals, called shelterin, consists of six core subunits: TRF1, TRF2, POT1 and TIN2, TPP1 and RAP1. This complex and its accessory factors are central players in the maintenance of genome integrity by shielding telomeric DNA from unwanted DNA repair activities. In yeast, telomeric protein complexes are different. While S. cerevisiae telomeres are bound by scRAP1, which interacts with RIF1 and RIF2, S. pombe features a telomeric complex with at least six subunits.
In both yeasts and humans, it has been observed that telomeres can be tethered to the nuclear periphery and exert a gene regulatory effect by forming a heterochromatic structure that reversibly suppresses the transcription of their nearby subtelomeric proximal genes. This telomere position effect (TPE) or telomeric silencing relies on epigenetic regulation by histone modifications.
Antigenic variation in trypanosomes
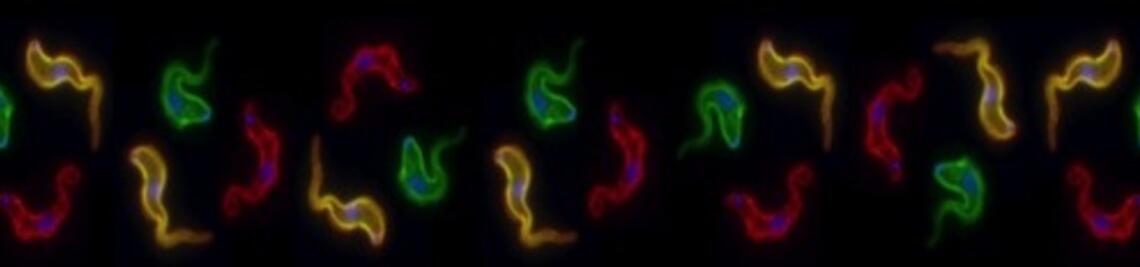
In microbial pathogens, virulence gene expression can be regulated by the telomere structure as those genes are often found adjacent to telomeres. In trypanosomes, the developmentally regulated subtelomeric variant surface glycoprotein (VSG) genes are the main virulence determinants in the mammalian-infectious bloodstream form (BSF). The cell surface of the BSF parasite is densely covered with a single species of VSG. To escape the mammalian host immune response, trypanosomes depend on antigenic variation of its VSG coat. Antigenic variation in trypanosomes is characterized by monoallelic transcription and by switching of the VSG gene. Key structures of antigenic variation are 15 specialized subtelomeric transcription units, the expression sites (ESs).
There is an intricate connection between the subtelomeric VSG genes and the telomeric proteins in trypanosomes. The telomere-binding proteins TbTRF and TbTIF2 suppress VSG switching. TbRAP1 promotes ES silencing by controlling the telomeric chromatin structure. Depletion of TbRAP1 leads to derepression of ES-linked VSG genes in BSF and PCF cells. In contrast to the yeast TPE mechanism, the TbRAP1-mediated VSG silencing in trypanosomes seems to act independently of Sir proteins, as depletion of the trypanosome Sir2 homolog does not prevent VSG silencing. The studies done on TbTRF, TbTIF2, and TbRAP1 provide information on how trypanosome telomeres maintain the inactive state of ESs and suppress recombination. If and how telomere-binding proteins contribute to ES silencing initiation has not yet been studied.
Telomeres & VSG expression
We recently used quantitative interactomics to determine the composition of telomere protein complexes in T. brucei BSF and PCF stages to learn more about the structure and functions of telomeres in trypanosomes. We identified several novel telomeric proteins, which are in the focus of our current research. Our data suggest a different telomere complex composition in the two forms of the parasite. One of the novel telomere-binding proteins, TelAP1, forms a complex with telomeric proteins TbTRF, TbRAP1 and TbTIF2 and influences ES silencing kinetics during developmental differentiation.
In collaboration with Dr. Falk Butter, head of the Quantitative Proteomics at the IMB in Mainz, we now want to investigate the impact of these novel telomere-associated protein complexes on VSG expression site regulation and structure in T. brucei. We want to know how these proteins bind telomeric DNA and what the composition of different telomere-associated complexes is. Furthermore, we want to learn if they are involved in important telomere-associated processes such as VSG regulation, recombination events or telomere maintenance.
Further reading:
TelAP1 links telomere complexes with developmental expression site silencing in African trypanosomes (2018) Reis H, Schwebs M, Dietz S, Janzen CJ, Butter F. Nucleic Acids Res. Jan 29. doi: 10.1093/nar/gky028